Five Pitfalls to Avoid When Generating Your Knockout Cell Line Knockout cell lines are an...
How Clonal Hematopoiesis Impacts Human Aging and Disease
How Clonal Hematopoiesis Impacts Human Aging and Disease
Clonal hematopoiesis refers to the expansion of a specific blood cell population harboring an acquired somatic mutation. The expansion of this advantageous clone results in a disproportionate population of blood cells with these selective mutations; as this cohort of cells expands, it will make up a larger portion of the overall blood cell population. Early studies suggested that 10-20% of the population over the age of 70 has clonal hematopoiesis. However, recent publications using advanced NGS technologies have reported this number may be much higher.
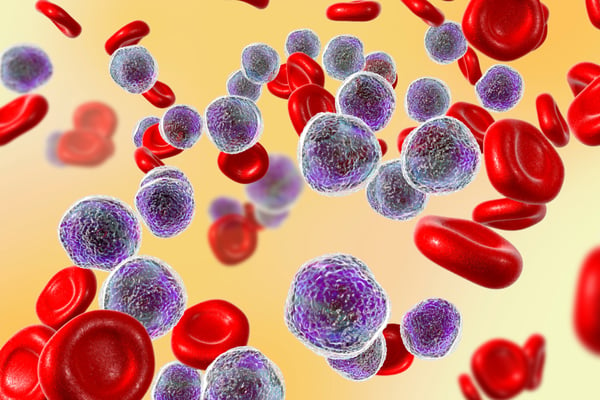
Clonal Hematopoiesis Origin and Detection
Clonal hematopoiesis is thought to originate from hematopoietic stem cells, which develop into all blood cell types. These stem cells, HSCs, are maintained for life and may acquire mutations in the protein-coding exon at a rate of one per decade. This number seems small, until you consider the elderly population which may be harboring multiple mutations in these all-important gene sequences resulting in genetic mosaicism, and potentially leading to myeloid malignancies and other diseases. Individuals who have been identified to have clonal hematopoiesis but are asymptomatic are referred to as having clonal hematopoiesis of indeterminate potential (CHIP). Individuals with CHIP may eventually develop a disorder or myeloid malignancy, but many do not.
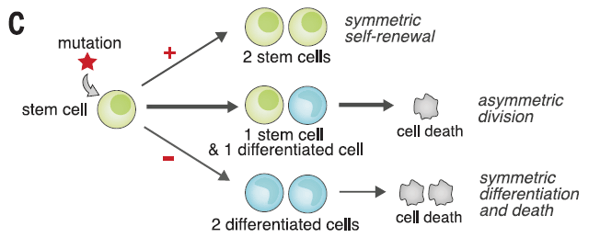
Clonal hematopoiesis is detected using DNA sequencing technologies that identify a mutation in blood or bone marrow samples, but this mutation is not present in other tissues. This is key because CHIP status can be associated with an increased risk of hematologic malignancy or cardiovascular disease. The ability to detect a clonal population depends on the sensitivity of the detection method and is reported as variant allele frequency (VAF; the proportion of mutated DNA at a given locus). This figure can be used to report on the approximate size of the clone population. Standard Next Generation Sequencing can detect a VAF of approximately 0.01 or 1%.
Use RareSeq to Identify Ultra-rare Gene Variants
Canopy’s proprietary RareSeq® technology is orders of magnitude more sensitive, pushing the limit of detection to 1:10,000 or a VAF of 0.0001 (0.01%). This enables the identification of ultra-rare gene variants and earlier diagnosis of clonal hematopoiesis. RareSeq uses the TruSight® Myeloid panel from Illumina, which is comprised of genes found to be commonly mutated in myeloid malignancies, as well as those specific to clonal hematopoiesis, to identify the CHIP status of a sample. The limit of detection here is important because, while the prevalence of clonal hematopoiesis increases with age, identifying it early may allow for earlier detection of deleterious mutations, better stratification of patients during clinical trials, and, ultimately, better patient outcomes.
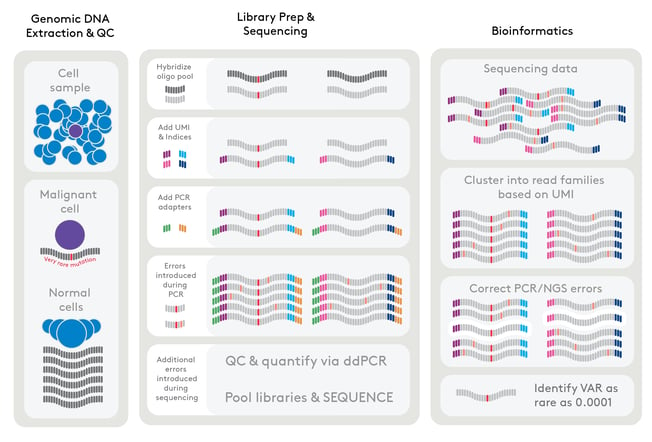
RareSeq becomes an important tool in the study of clonal hematopoiesis when thinking about population genetics. As noted previously, early estimates reported that 10-20% of the population over 70 years of age have evidence of clonal hematopoiesis. But previous methods did not allow for the detection at 1:10,000, missing variant clonal mutants below this threshold. While these early studies found that CHIP was virtually absent in populations under 40 years of age and indicated a sharp increase after 60, as detection methods became more sensitive, studies found that younger individuals, even those sub-40 years of age, are harboring potentially pathogenic mutations. One such study from 2016 applied RareSeq’s methodology to a cohort of healthy 50-60 year olds. They surprisingly found that 95% of the individuals studied harbored mutations at DNMT3S and TET2.
Another impact of CHIP status is eligibility as a donor for hematopoietic stem cell transplantation (HSCT). A study out of Washington University in St. Louis found that the assumption of negative CHIP status in bone marrow donors younger than 40 years of age was incorrect. Clonal mutations were discovered in 44% of the donor population (aged 20-58, median age of 28) and all of these were found to be engrafted in the recipient’s blood cell population. Strikingly, >80% of the mutations were predicted to impart functionally meaningful impacts on the affected gene product. The study utilized RareSeq® and called for further research into screening methods of bone marrow donors.
Clonal hematopoiesis has been shown to impact larger percentages of the population than initially thought, and is an indicator of myeloid and hematological malignancies, among other diseases. Having a technology that can assess CHIP status of a patient sample is a significant tool in several key areas of human health such as population genetics and stem cell donor screening. As research into this area continues, RareSeq will become a powerful tool in assessing clonal hematopoiesis.
RareSeq is a Complete Service for Error-Corrected Sequencing.
References:
1. Welch JS, Ley TJ, Link DC, et al. The origin and evolution of mutations in acute myeloid leukemia. Cell. 2012;150(2):264-278. doi:10.1016/j.cell.2012.06.023
2. Steensma DP, Bejar R, Jaiswal S, et al. Clonal hematopoiesis of indeterminate potential and its distinction from myelodysplastic syndromes. Blood. 2015;126(1):9-16. doi:10.1182/blood-2015-03-631747
3. Malcovati L, Cazzola M. The shadowlands of MDS: idiopathic cytopenias of undetermined significance (ICUS) and clonal hematopoiesis of indeterminate potential (CHIP). Hematology Am Soc Hematol Educ Program. 2015;2015:299-307. doi:10.1182/asheducation-2015.1.299
4. Young AL, Tong RS, Birmann BM, Druley TE. Clonal hematopoiesis and risk of acute myeloid leukemia. Haematologica. 2019;104(12):2410-2417. doi:10.3324/haematol.2018.215269
5. Wong WH, Bhatt S, Trinkaus K, et al. Engraftment of rare, pathogenic donor hematopoietic mutations in unrelated hematopoietic stem cell transplantation. Sci Transl Med. 2020;12(526):eaax6249. doi:10.1126/scitranslmed.aax6249
6. Young AL, Challen GA, Birmann BM, Druley TE. Clonal haematopoiesis harbouring AML-associated mutations is ubiquitous in healthy adults. Nat Commun. 2016;7:12484. Published 2016 Aug 22. doi:10.1038/ncomms12484
7. Watson CJ, Papula AL, Poon GYP, et al. The evolutionary dynamics and fitness landscape of clonal hematopoiesis. Science. 2020;367(6485):1449-1454. doi:10.1126/science.aay9333