Five Pitfalls to Avoid When Generating Your Knockout Cell Line Knockout cell lines are an...
Skip the Plasmids: Why You Should Use RNPs for Your CRISPR Projects
Skip the Plasmids: Why You Should Use RNPs for Your CRISPR Projects

CRISPR-Cas9 has revolutionized gene editing, providing a cost-effective, efficient, and relatively facile system to produce gene knockouts, point mutations, tags, and other mutations in cell lines and animal models. The CRISPR-Cas9 system relies on a guide RNA (gRNA) for targeting and Cas9, an endonuclease. Working together, the two components generate a targeted double-strand break that is critical to the gene editing process (figure 1). In the cell, the gRNA is active as RNA and Cas9 as protein—no matter the delivery method, they must be transcribed/translated in order to be active.
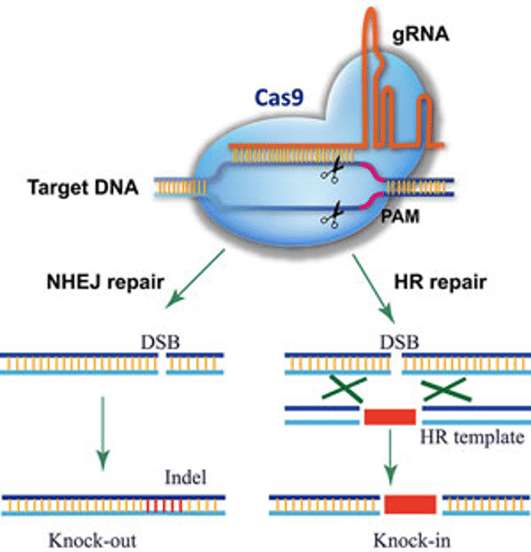
Figure 1:The CRISPR-Cas9 system generates a targeted double-strand break that can be harnessed for gene editing. Image credit: Ding Y, Li H, Chen L-L and Xie K (2016) Recent Advances in Genome Editing Using CRISPR/Cas9. Front. Plant Sci. 7:703. doi: 10.3389/fpls.2016.00703
Plasmid delivery of CRISPR reagents, and in particular “all-in-One” plasmids that contain all of the CRISPR-Cas9 components—sgRNA and Cas9—on a single plasmid, have been an attractive choice due to their simplicity. However, these plasmids can present challenges researchers may not expect.
Challenge 1: Cas9 must be transcribed and translated from plasmids
Cas9 is active in the cell as protein. Conversely, sgRNAs (required for Cas9 targeting) are active as RNA. As a result, there can be a mismatch in timing of activity of the two components when expressed off the same plasmid as Cas9 must be further translated to protein (figure 2). This can result in a reduction of efficiency of Cas9 and reduced editing activity.
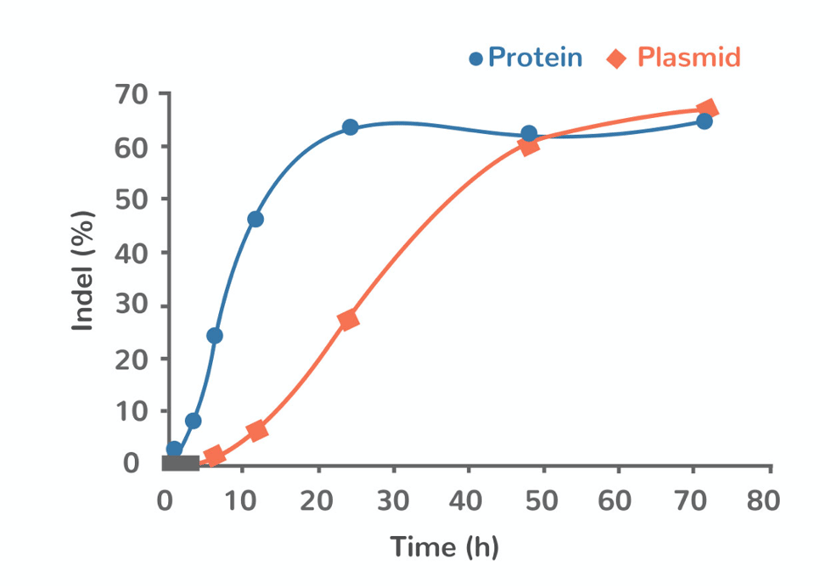 Figure 2: Time-course of Cas9 activity following delivery into the cell in either plasmid format (orange) or as protein (blue). Adapted from Sojung Kim et al. Genome Res. 2014;24:1012-1019
Figure 2: Time-course of Cas9 activity following delivery into the cell in either plasmid format (orange) or as protein (blue). Adapted from Sojung Kim et al. Genome Res. 2014;24:1012-1019
Challenge 2: Plasmids are active for too long
While it can take some time to get active Cas9 in the cell via plasmid delivery, expression levels can remain appreciable for several days, and even longer if the plasmid integrates into the genome (figure 3). Prolonged expression of Cas9 increases the probability of off target effects—a known concern with the CRISPR-Cas9 system in general (figure 4). Off target mitigation strategies look to limit the time that Cas9 is active—ideally, Cas9 should make its targeted cuts and then quickly be degraded.
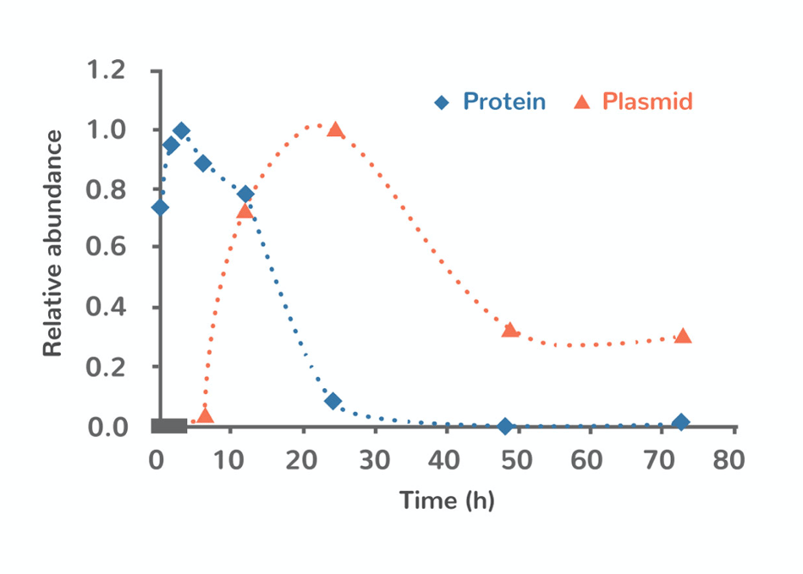 Figure 3: Degradation profiles of Cas9 delivered as plasmid (orange) or protein (blue). Adapted from Sojung Kim et al. Genome Res. 2014;24:1012-1019
Figure 3: Degradation profiles of Cas9 delivered as plasmid (orange) or protein (blue). Adapted from Sojung Kim et al. Genome Res. 2014;24:1012-1019
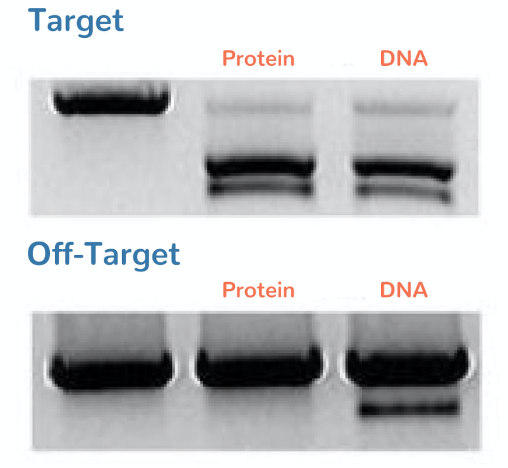
Figure 4: T7E1 assay demonstrating cutting efficiency at target and off target sites. Adapted from Sojung Kim et al. Genome Res. 2014;24:1012-1019
Challenge 3: Plasmids may integrate into the genome
After delivery to the cell, plasmids may integrate randomly into the genome. While constitutive activity of CRISPR-Cas9 components are problematic (as discussed above), there is an additional concern. Integration into the host cell genome is random and unpredictable. Integration into and subsequent disruption of other genes may produce unwanted off target effects that may convolute data interpretation.
Fortunately, an efficient solution exists to combat these challenges. Cas9 is now commercially available in protein form. Cas9 protein can be complexed with gRNAs prior to delivery into the cell. The resulting ribonucleoprotein (RNP) is then directly transfected or nucleofected into cells, where the reagents are immediately active and then quickly degraded. The result is a reagent format that is efficient and largely free of off target activity.
Want more efficient gene editing with fewer off-target effects?
Check out our Purified Cas9 Protein.
Need gRNAs custom-designed for your project along with your Cas9 protein?
Check out our CRISPR Complete Kits.
Have a gene, mutation, and cell line but don't want to tackle gene editing yourself?
Our Cell Line Editing Service is the perfect match.